This resource is available for free download
Darren collected a soil sample from Drumpellier Park in Coatbridge, North Lanarkshire. He extracted the DNA from the soil sample and sent it for sequencing.

Where do we find DNA? DNA is found in the nucleus of most cells. It contains the instructions for the development and function of living things – this is called the genetic code. The soil contains DNA from living things (and things that once lived). Every living thing has its own DNA fingerprint. We can use this fingerprint as a clue to identify it. Non-living things do not have any DNA.
The sequencing results contained DNA sequences for each of the living things in his soil sample. This is the DNA sequence for one type of bacteria in the soil:
NNNTTANCATGCANGTCGAACGATGAAGCCCTTCGGGGTGGATTAGTGGCGAACGGGTGAGTAACACGTGGGCAATCTGCCCTGCACTCTGGGACAAGCCCTGGAAACGGGGTCTAATACCGGATACTGATCGCCTTGGGCATCCTTGGTGATCGAAAGCTCCGGCGGTGCAGGATGAGCCCGCGGCCTATCAGCTTGTTGGTGAGGTAATGGCTCACCAAGGCGACGACGGGTAGCCGGCCTGAGAGGGCGACCGGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGGAGGCAGCAGTGGGGAATATTGCACAATGGGCGAAAGCCTGATGCAGCGACGCCGCGTGAGGGATGACGGCCTTCGGGTTGTAAACCTCTTTCAGCAGGGAAGAAGCGAAAGTGACGGTACCTGCAGAAGAAGCGCCGGCTAACTACGTGCCAGCAGCCGCGGTAATACGTAGGGCGCGAGCGTTGTCCGGAATTATTGGGCGTAAAGAGCTCGTAGGCGGCTTGTCGCGTCGGTTGTGAAAGCCCGGGGCTTAACCCCGGGTCTGCAGTCGATACGGGCAGGCTAGAGTTCGGTAGGGGAGATCGGAATTCCTGGTGTAGCGGTGAAATGCGCAGATATCAGGAGGAACACCGGTGGCGAAGGCGGATCTCTGGGCCGATACTGACGCTGAGGAGCGAAAGCGTGGGGAGCGAACAGGATTAGATACCCTGGTAGTCCACGCCGTAAACGGTGGGCACTAGGTGTGGGCGACATTCCACGTCGTCCGTGCCGCAGCTAACGCATTAAGTGCCCCGCCTGGGGAGTACGGCCGCAAGGCTAAAACTCAAAGGAATTGACGGGGGCCCGCACAAGCGGCGGAGCATGTGGCTTAATTCGACGCAACGCGAAGAACCTTACCAAGGCTTGACATACACCGGAAACGTCCAGAGATGGGCGCCCCCTTGTGGTCGGTGTACAGGTGGTGCATGGCTGTCGTCAGCTCGTGTCGTGNNNTGTTGGGTTAAGTCCCGCAACGAGCGCAACCCTTGTCCCGNNTTGCCAGCAGGCCCTTNNNN

So can you identify this bacteria? Yes! If you compare this DNA sequence to a great big worldwide database of DNA sequences to see how closely it matches the DNA of an organism that scientists have already identified. It is a bit like using a computer database to compare one sentence to all the books in a library to find out which book it came from. There is a tool that we use for this called BLAST.
Click here to open the National Center for Biotechnology Information BLAST tools in a separate window.
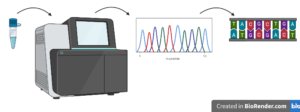
DNA is made of building blocks called nucleotides. Nucleotides are chemical molecules. The nucleotides that build DNA are called deoxyribonucloetides. That’s a long word!

Look back at the DNA sequence. You can see it is a string of the letters A, G, C and T. These letters are a shorter way of writing the names of the nucleotide molecules adenine (A), guanine (G), cytosine (C) and thymine (T).
When Darren sent his soil sample for DNA sequencing, this process worked out the order of the nucleotides in the strands of DNA extracted from his soil. [The N’s in his sequence stand for unknown bits of DNA in this strand that couldn’t be identified].
You are going to do a nucleotide BLAST, so click on “Nucleotide BLAST” on the NCBI page.

Don’t worry if the nBLAST page looks a bit complicated. Let’s work through it step by step.
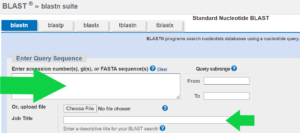
Copy and paste the DNA sequence into the top box (big green arrow). You can put a name for your search into the Job Title box (little green arrow) if you want e.g. Darren’s bacteria search.
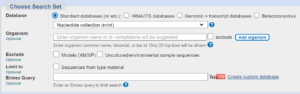
You don’t need to change any settings or add any extra information in the second box to search the standard database of nucleotides.

In the last box, choose to look for somewhat similar sequences using a computer algorithm called blastn.

Click on BLAST to blast off with your search!
You will get a page that shows the status of your search. When lots of people are searching the database at the same time, it may take a little while.

The results window shows the hits in the database:
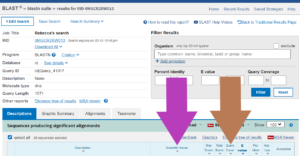
You need two bits of information to identify what living thing your DNA sequence comes from: the scientific name of any organisms that have DNA sequences that match your sequence (purple arrow) and how close the match is (query coverage, shown by brown arrow).
Scientific names are in Latin so that scientists from all over the world know they are talking about the same organism. The query coverage is a percentage (%) and tells you how much your DNA sequence overlaps with the sequences in the database. The higher the percentage, the more likely it is that your DNA came from this organism.
What bacteria did Darren find?

Want to find out more about DNA and sequencing?
Sanger Sequencing | Let’s Talk Science (letstalkscience.ca)
The DNA Code with Alison Woollard. Clip from the Royal Institution “Life Fantastic” Christmas Lecture series
